TubeTK/Events/2010.10.19
From KitwarePublic
< TubeTK | Events
Jump to navigationJump to search
Revision as of 16:51, 19 October 2010 by Danielle.pace (talk | contribs)
- Dashboard: https://www.kitware.com/CDash/index.php?project=TubeTK
- Registration re-write needed
- Wiki Documentation: http://public.kitware.com/Wiki/TubeTK
Stephen
Completed
- Ridge extractor success
- Email requests for TubeTK
- Licensing changes
- Blur3D -> Blur
Ongoing
- Radius extractor tests
- 2D and 3D only :(
- Funding re-allocation
Upcoming
- Application in plant biology
- Proposal with Paul / Liz
Romain
- Implementation on the ComputeTortuosity module (only Distance Metric)
- Started debugging
- Future
- More debugging! ---> hopefully testing soon
- #tube command
- Implementation of the other 2 methods : ICM SOAM
Danielle
Sliding organ registration
- Fixed "duh" problem from Thursday - extracting tangential and normal components before computing regularization
- Regularization smoothing in normal direction completed
- Added intensity distance function (mean squares)
XCAT phantom
- Got XCAT running on my machine (thanks Andinet!)
- Learned XCAT parameters
- Wrote isbiValidation.py:
- --basic (XCAT mode 0 - phantom mode): with respiratory period 5s and cardiac period 1s, generates two images at 0s and 2.5 s, saves as .mhd
- --gold (XCAT mode 2 - spherical lesion generation mode): creates gold standard motion field, and saves as .txt (index, vector in 6 columns). Then run itkImageToImageDiffusiveDeformableRegistrationGenerateGoldStandardTest to create itk::Image<VectorType, Dimension> .mhd for visualization
- XCAT phantom gives gold standard vectors for surface borders (XCAT mode 4), but no internal vectors! So...
- for index in range - generate spherical lesion at index, use nurbs_save = 1 to save lesion.nrb, then calculate centroid of lesion control points
- --segmentation (XCAT mode 5 - save anatomical variation mode, or run XCAT mode 0 with save_nurbs = 1) to save surfaces (.nrb text file). Then create new .txt file for the surface you want. Then run itkImageToImageDiffusiveDeformableRegistrationGenerateNormalsTest to create itk::Image<PixelType, Dimension> of control points
- Unfortunately: (1) control points need interpolation, can't find NURBS support in VTK; (2) XCAT has MANY types of surfaces for one organ - ex all intracardiac surfaces, etc, and it would be a lot of work to sort through them; (3) Perfect segmentation not great for registration method evaluation - removes segmentation error component of total error.
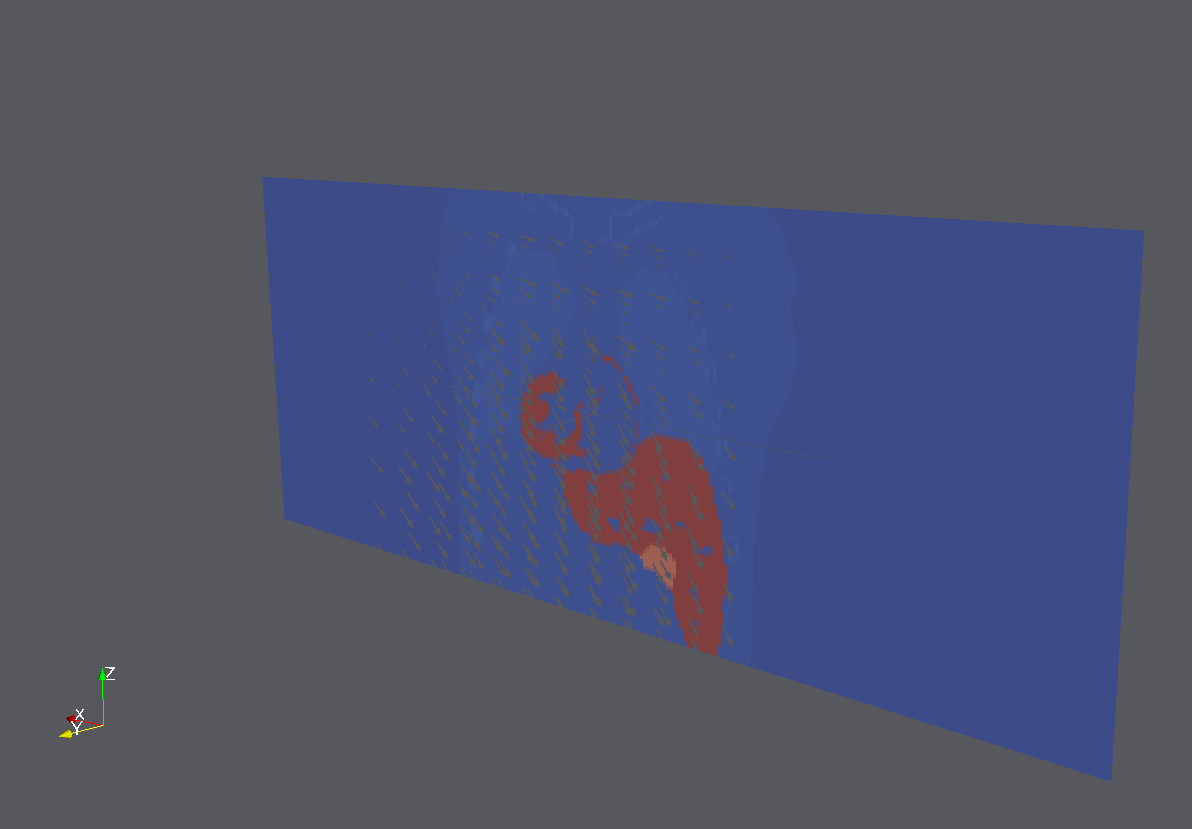
- Ex: right lung
- --segmentation (XCAT mode 5 - save anatomical variation mode, or run XCAT mode 0 with save_nurbs = 1) to save surfaces (.nrb text file). Then create new .txt file for the surface you want. Then run itkImageToImageDiffusiveDeformableRegistrationGenerateNormalsTest to create itk::Image<PixelType, Dimension> of control points
Talking points
- Underlying XCAT deformation fields too perfect? They are likely interpolating the border vectors.
Future
- Boundary conditions
- Compute weighting term = distance to organ border
- Extract organ border normals from input data
- Run registration + compare to gold standard for validation #s